Comprehensive analysis of all genetic material in complex microbial communities using advanced whole-genome shotgun sequencing. Culture-independent, high-throughput, high-resolution — powered by NGS and AI-enhanced analytics.
Metagenomics is the direct study of the collective genetic material DNA recovered from an environmental or biological sample, without the need to culture individual organisms in a lab. Unlike traditional microbiology, which examines one microbe at a time, metagenomics captures the entire community of bacteria, archaea, fungi, viruses, and other microbes in a single sequencing run.
This approach is transforming our understanding of microbial ecosystems in the human gut, soil, water, and beyond, revealing who is there, what they are doing, and how they interact, all with unprecedented resolution and speed.
MGBIO drives innovation in Tech-Bio, offering tailored service packages, high-throughput data analytics, and web-based solutions for biological and health-related big data. We focus on next-generation tools, protocols, and products to enhance diagnostics, improve patient outcomes, and address region-specific challenges.
Based at the National Science & Technology Park (NSTP), NUST, Islamabad, MGBio is a leading Tech-Bio company offering genomic services, AI/ML model development, bioinformatics solutions, proteomics, molecular biology services, scientific content drafting, and professional training programs.
Choose from three complementary sequencing approaches, each tailored to specific microbial targets and research objectives.
Targets conserved regions of the 16S rRNA gene to profile bacterial and archaeal communities. This method provides efficient, cost-effective taxonomic identification, typically at the genus to species level
Focuses on the highly variable Internal Transcribed Spacer (ITS) regions, particularly ITS2, for accurate identification of fungal species. It enables discrimination between closely related fungal taxa in complex samples
Sequences all genomic DNA present in a sample, providing comprehensive coverage of bacteria, archaea, fungi, viruses, and other microbes. This approach allows high-resolution analysis at species and strain level
Choose your platform based on read length, budget, and target. We also analyze datasets from any external vendor
16S
✓ Recommended
18S
✓ Recommended
WGS
✓ Recommended
16S
✓ Recommended
18S
✓ Recommended
WGS
✓ Recommended
16S
~ Conditional
18S
~ Conditional
WGS
✓ Recommended
16S
✓ Recommended
18S
✓ Recommended
WGS
✓ Recommended
16S
✗ Not Rec.
18S
✗ Not Rec.
WGS
✗ Not Rec.
Ensure your DNA samples meet these thresholds for optimal sequencing results.
Min. DNA quantity
( Rec. > 400ng)
Min. Concentration
Min. Volume
High Molecular Weight intact DNA required
Comprehensive analysis pipelines with three tiers of output reports. Essential, Beneficial, and Extraordinary, with integrated AI capabilities.
Microbes TargeteD
Bacteria & Archaea
AI Capabilities
AI Capabilities
AI Capabilities
Microbes Targeted
Fungi
AI Capabilities
BENEFICIAL
AI Capabilities
AI Capabilities
All Micro-organisms
Microbes Targeted
AI Capabilities
AI Capabilities
AI Capabilities
Every project comes with a comprehensive set of outputs, designed to support your research from data generation through scientific publication
Comprehensive summary of the methodologies applied, including sequencing workflow, quality control steps, and analytical approaches used throughout the project.
High-throughput sequencing files delivered via secure server access, accompanied by detailed QC reports including demultiplexing and pre-processing statistics.
Full QC metrics plus raw FASTQ files and a consolidated quality control summary report covering all sequencing runs.
Processed results including assembly, binning, taxonomic classification, and functional annotation, all presented as publication-ready figures.
Up to five 60-minute consultation sessions with project specialists to assist with data interpretation, manuscript support, and editorial processes.
MGBio provides scientific writing services including manuscript drafting, grant proposals, and technical reports as an integral component of broader offerings
Our experts are ready to design a custom metagenomics project for your exact research needs. First consultation is free.
Real publication-quality figures generated by MGBio across all analysis tiers. Every project delivers outputs at this standard
MAG recovery distributions, species abundance quantiles, and community-level performance comparison
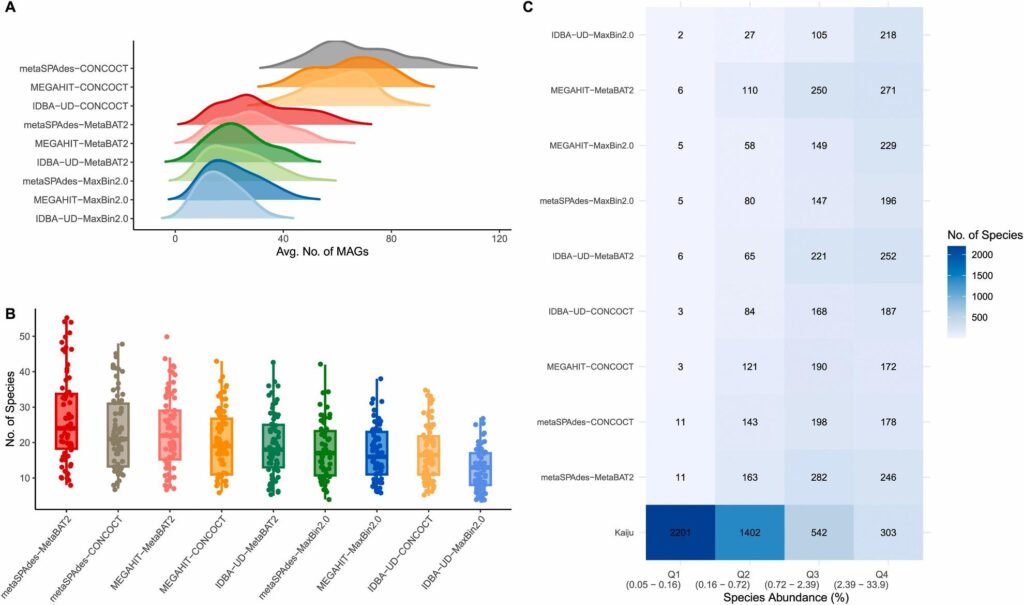
Strain counts per tool, species-level strain bubble plots, and strain quality heatmaps across abundance classes
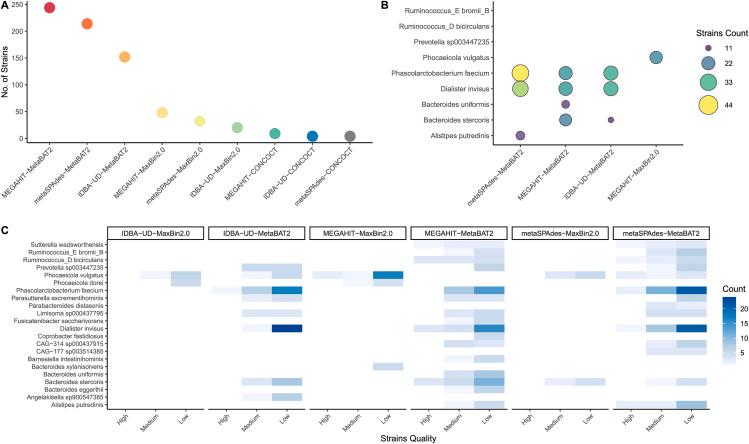
Species recovery status heatmap across tools and abundance-stratified recovery proportion analysis
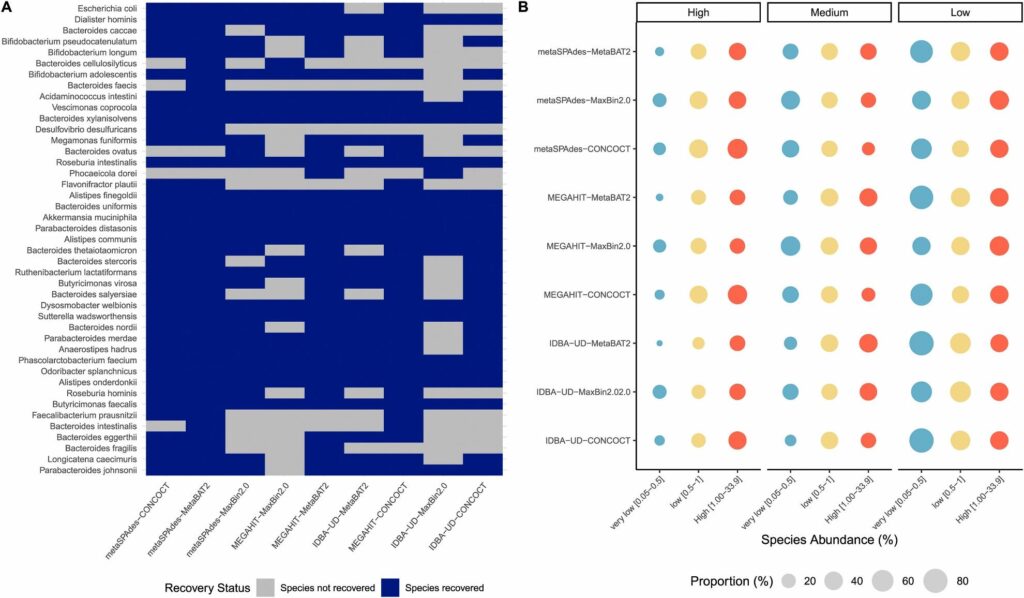
Our team’s published research underpins the analytical rigor and innovation behind our services.
2025
Qayyum H, Talib MS, Ali A, Kayani MUR. Evaluating the potential of assembler-binner combinations in recovering low-abundance and strain-resolved genomes from human metagenomes. Heliyon. 2025 Jan 14;11(2):e41938. doi: 10.1016/j.heliyon.2025.e41938.
2025
Qayyum H, Ishaq Z, Ali A, Kayani MUR, & Huang L. Genome-resolved metagenomics from short-read sequencing data in the era of artificial intelligence. Functional & Integrative Genomics, 25(124). doi: 10.1007/s10142-025-01625-x
2026
Qayyum H, Raziq MF, Manzoor H, Zaidi SSA, Ali A, Kayani MUR. Efficient De Novo Assembly and Recovery of Microbial Genomes from Complex Metagenomes Using a Reduced Set of k-mers. Interdisciplinary Sciences. 2026 Mar;18(1):151–164. doi: 10.1007/s12539-025-00722-6.
2026
Nasir MM, Qayyum H, Shuhui S, Ali A, Kayani MUR. MetaBolt: A computationally efficient pipeline for the rapid recovery of metagenome-assembled genomes. Computational Biology and Chemistry. 2026 Mar 11;123:109006. doi: 10.1016/j.compbiolchem.2026.109006.
Development & Training of AI & ML based models for efficient predictions & data-driven analyses for better & informed decisions making
Analysis Suites & High Throughput Software Pipelines and Packages / Standalone & web based packages / Real-Time & Knowledgebase Databases
DNA/RNA isolation & cDNA synthesis. PCR & Real-time PCR (Probe based/SYBR Green). Cloning & Recombinant DNA technology
Protein Isolation & Mutagenesis / SDS PAGE / Western Blotting / ELISA services / Mass Spectrometry / HPLC & FPLC / Bioinformatics Analysis of Proteomes
Editorial Services, Blog Writing. Manuscript Drafting. Language editing. Graphical Abstracts, Quality Images, Figures and illustrations.
Curriculum and program development, consultancy services to govt. & private sectors, projects & guidance, institutional framework and policy development.
Trainings, Lectures, Seminars and Hands-on Workshops as per Customer Feedbacks and Demands (Online & in person). Internships and Guidance.
Genome sequencing, Assembly & Annotation Services / Bioinformatics analysis, Metagenomics & Transcriptomics Data Analysis / Comparative Genomics
Chat with us